
How to compute ΔF/F from calcium imaging data
By Dr Peter Rupprecht, University of Zurich
Key takeaways
- ∆F/F reflects intracellular calcium levels, but often in a nonlinear way
- ∆F/F was initially introduced for organic dye calcium indicators in the 1980s
- GCaMP behaves differently due to its low baseline brightness and its nonlinearity
- There is no single recipe on how to compute ∆F/F
- Computation of ∆F/F must be adapted to cell types, activity patterns, and noise
- Interpretation of ∆F/F requires knowledge about indicators, cell types, and confounds
- Potential confounds are brain motion, neuropil contamination, and response variability across neurons
What is calcium imaging?
Calcium imaging is an amazing technique. With calcium imaging, we can record the activity of neurons in living animals and see these neurons while doing so. The mesmerizing beauty of these movies can make us fall in love with this technique immediately. However, from a scientific perspective, the goal is not just to capture beautiful pictures of blinking neurons but to extract meaning from them and interpret the results – and this takes a couple of steps and some background knowledge.
One of these processing steps is to compute the normalized fluorescence, ∆F/F. To do so, you take the fluorescence trace over time, F(t), and compute (F-F0)/F0, abbreviated as ∆F/F, with the baseline fluorescence F0. However, every paper seems to use slightly different procedures to compute F0 and ∆F/F. What is the most solid and scientifically sound way to do so? More generally, why do we use this normalization procedure to begin with? And, finally, how should we interpret ∆F/F in the end?
Calcium imaging video recorded in hippocampal CA1 pyramidal cells of an awake mouse by Sian Duss and Peter Rupprecht.
The historical origins of ∆F/F0
Why we use ∆F/F0 for normalization
Let’s start with the simplest question: why do we need to normalize the fluorescence changes to start with? There is a simple answer: because some neurons appear brighter than others, and if we didn’t normalize, fluorescence changes computed for bright cells would be larger than for dim cells.
Okay. But why can’t we just normalize by the standard deviation of the recorded trace (as does the z-score), or by the mean fluorescence? Why do we need to use the baseline F0? This question can be best answered when looking back at the history of calcium imaging. Historically, ∆F/F has been related directly to changes of the calcium concentration. The baseline F0 represents the fluorescence value in the presence of the calcium concentration that is observed for the cell ideally in the absence of activity, and any increase of F(t) below this zero level indicates a change of calcium compared to this baseline level.
The linear and nonlinear regimes of ∆F/F0
However, ∆F/F is not simply a linear function of the calcium concentration. In early papers on calcium imaging in the 1980s, expressions for [Ca2+] as a function of ∆F/F were given for the first time. This equation is based on the assumption of a chemical equilibrium between calcium ions and molecules, each of which binds a single calcium ion. Later work reshaped the relationship into now more well-known forms:
This equation represents a saturating relationship. It is approximately linear for small calcium concentrations. “Small” means small compared to the dissociation constant of the calcium indicator, kD. There are three key takeaways from this equation and its assumptions: First, all calcium indicators saturate for high calcium concentrations, , which can be induced by high spike rates. There is no way around this limitation. Second, these calcium indicators are linear only for low calcium concentrations, . Therefore, the calcium concentration range, which varies from cell to cell, and the dissociation constant of the calcium indicator together determine the linear and saturating regime for each cell. Third, the equation derives from the assumption of a reaction equilibrium. For fast events such as calcium influx during an action potential, the reaction equilibrium is not reached during the action potential, and the equation becomes therefore less accurate and more heuristic. Fourth, this equation was derived and analyzed at a time when genetically encoded calcium indicators like GCaMP were still in their infancy. The specifics of GCaMP-like sensors were not considered.
In general, despite the limitations, the approximation that ∆F/F is proportional to calcium concentration changes is useful. It is what makes ∆F/F a more meaningful metric than a simple z-score or raw fluorescence values.
∆F/F0 in the era of GCaMP
GCaMP binds four calcium ions
The above equations for ∆F/F were based on models that assumed the binding of a single calcium ion to a single calcium indicator. With the advent of genetically encoded calcium indicators, in particular GCaMP, this assumption was not valid any more. Why? GCaMP differs because it has four binding sites for calcium ions. The binding of these sites is not independent, a phenomenon termed cooperative binding. The formalism to describe such a cooperative binding existed already in the form of the Hill equation (you may have seen this equation when studying the cooperative binding of oxygen to hemoglobin!), which is effectively a generalization of the above equation:
with the number of binding sites, . In the case of n=1, we arrive at the same formula as above. GCaMP has 4 binding sites, but since they are not fully cooperative, the effective (and therefore the nonlinearity) is lower and ranges between 2-3 for GCaMP indicators (e.g., nGCaMP6f ≈ 2.3, nGCaMP7f ≈ 3.1, nGCaMP8s ≈ 2.2).
A sigmoid transfer curve for GCaMP
As a result, the calcium-to-fluorescence relationship is now a sigmoid, being nonlinear both for and for , with an approximately linear and highly sensitive working range around . This sigmoid relationship can in practice be seen as a contrast enhancer and is similarly used in image processing software like Photoshop or GIMP. Maybe early GCaMP sensors were so successful not only because they were great work of engineering but also because they largely enhanced the contrast of our calcium images, with sharp and striking transitions between low and high levels of activity, resulting in the beautiful blinking of neurons.

Non-linear sigmoidal relationship of calcium to fluorescence typical for e.g. GCaMP6f. kD is the dissociation constant of the calcium indicator. The sigmoidal transformation corresponds to a contrast enhancement as is commonly used for image processing: middle and right, picture taken in Lenzburg, Switzerland.
Depending on how the typical calcium concentration of a neuron overlaps with the sigmoidal curve centered around kD, the sensor will show different limitations. If the kD of such a cooperative calcium sensor is too low, it will saturate quickly for large numbers of spikes. However, if the kD of such a sensor is too high, it will not react at all towards single action potentials. Resting calcium levels of neurons are on the order of 50-200 nM, but may vary across cells and cell types. Calcium indicators, in turn, come with different kD values that make the calcium indicators behave more or less linear in the calcium range typically encountered in neurons: kD,GCaMP6f ≈ 375 nM, kD,GCaMP7f ≈ 150 nM, kD,GCaMP8s ≈ 46 nM. Therefore, to judge the linearity of a calcium sensor, one cannot rely only on the Hill coefficient but needs to also consider kD and how it relates to the range of calcium concentrations in the neuron of interest.
Low baseline brightness of GCaMP
An additional problem of calcium indicators such as GCaMP is the very low brightness during baseline (that is, with calcium at resting level). This property also comes with a big advantage: it reduces the background fluorescence from silent neuronal somata or dendritic trees and therefore makes activity more visible. However, at the same time, computing the value for F0 to obtain quantitative results becomes challenging. Even a small mistake in the determination of background fluorescence may have a big impact. And such mistakes are likely to occur, due to PMT offsets or neuropil contamination, etc. (see below!). Therefore, it is tricky to accurately determine F0, which in turn makes it difficult to quantitatively interpret ∆F/F values.
Does ∆F/F0 make any sense for GCaMP?
So, why should we use ∆F/F for GCaMP, despite these limitations? The main reason why we use it is indeed historical: it made sense for the previously used calcium indicators like Fluo-4 or OGB-1, and we just keep doing it. Second, we need some sort of normalization, otherwise bright neurons would always have large responses and dim neurons always weak responses. If you used z-scoring instead, highly active cells would be quenched, while silent cells would be artificially amplified. ∆F/F also seems intuitive because it is a positive metric of activity and zero when there is no activity (at least for neurons with sparse activity patterns). Third, newer GCaMP variants (GCaMP8s and GCaMP8m) seem to behave more linearly, which makes the relationship of ∆F/F to calcium again more interpretable. Therefore, ∆F/F is still or again useful (also see below for “How to interpret ∆F/F”).
How to compute ∆F/F0
F0 may be confounded by background
Computing ∆F/F is in principle very simple, as the formula ∆F/F suggests. So, how can we, in principle, compute F0 for a raw extracted fluorescence trace? F0 is the baseline fluorescence of the neuron when it is in the state of resting calcium concentration. This baseline fluorescence may, however, be confounded by background factors, and these background values should be subtracted from the recording before ∆F/F is computed. Don’t make the mistake to confound baseline with background – these are two distinct concepts.
As a first potential source of background, there can be an offset of the recording or digitization device. For example, the popular software ScanImage allows to subtract a “PMT offset”, which is supposed to be the intensity value when the PMT is switched on but not detecting any fluorescence; however, the software does not enforce the activation of this feature, and it can easily happen that an artificial negative or positive PMT offset is added to the entire image. In this case, efforts should be made to estimate the PMT offset after data acquisition and to subtract it from the raw data. Similar considerations apply to calcium imaging with other softwares or with cameras.
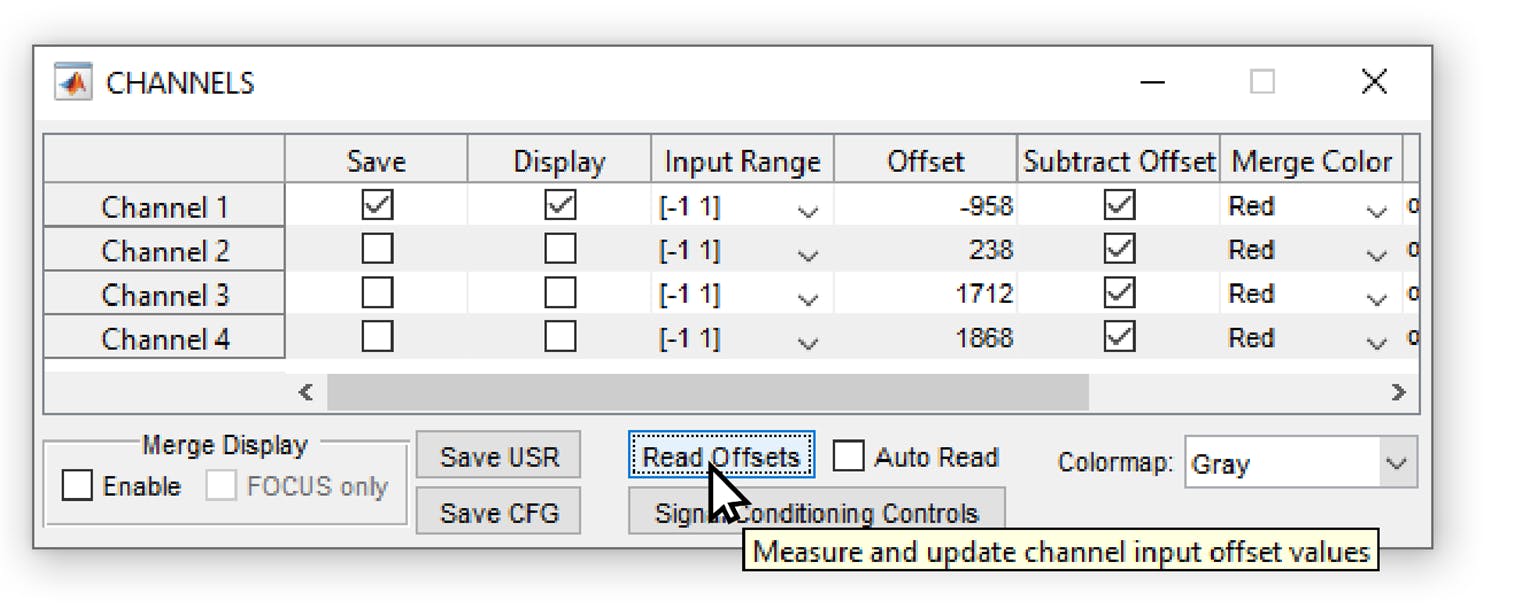
Screenshot from the software Scanimage (Vidrio Technology), commonly used to record two-photon microscopy data. If the PMT offset is not subtracted or not measured (using the “Read Offsets” button), there will be an arbitrary offset in the data as an instrument background.
Second, fluorescence from other cells can confound the baseline measurement of the cell of interest. For example, the diffuse dendritic arborizations which are not in focus but which still contribute a diffuse and significant fluorescence to each pixel, may increase the background and the estimated fluorescence value F0. For GCaMP with non-nuclear expression, this effect cannot be fully avoid for dense expression in a population of neurons. Related to that, toolboxes such as Suite2p perform a neuropil subtraction to compensate for this neuropil background. However, it can happen that neuropil subtraction is too aggressive and subtracts too much, creating a bias in the opposite direction. Other algorithms for cell extraction such as CaImAn and EXTRACT subtract background patterns before performing source extraction. Again, if this global background subtraction is too aggressive, it will also subtract the true baseline fluorescence from neurons, resulting in underestimates of F0.
These factors – PMT offset, neuropil contamination, and global background removal – need to be adjusted if needed or at least be kept in mind. In general, it is better to stay on the safe side to avoid division by zero. For example, one can add a safety margin chosen by experience, e.g., of 10 or 20 units (or whatever makes sense for your data), to the measured F0 value in order to prevent potential division by (close to) zero and the amplification of noise, or even sign inversion in case of negative values fo F0. While the arbitrary addition to F0 is difficult to justify based on a principled approach, it can be a practical solution to the problems that come with the low baseline brightness of GCaMP sensors.
Computing F0 from the median
Next, how can we calculcate F0 across an entire recording? Theoretically, in the absence of noise, the lowest fluorescence value across the entire recording should be taken as F0. However, one also needs to consider the effect of noise and potential drift. There are multiple ways to account for these potential problems. The simplest solution is to compute the median of a raw fluorescence trace (“fluo_trace”):
F0 = nanmedian(fluo_trace) - Matlab
F0 = np.nanmedian(fluo_trace) - Python
The median as a so-called “robust” metric is preferable compared to the mean since the distribution of fluorescence values typically is a skewed distribution, with periods of activity as positive outliers.
Computing F0 from the percentiles
The approach based on medians can be generalized and refined using percentiles:
F0 = nanmedian(fluo_trace) - Matlab
F0 = np.nanmedian(fluo_trace) - Python
Using a percentile of e.g. 20% can make sense when a lot of ongoing activity would shift the median to an F0 value that is clearly above the true baseline fluorescence. From this, it is already clear that there is no unique value of the percentile that is ideal for all recordings. For very low activity, a percentile of 50% is adequate (equivalent to taking the median), while the percentile should be lowered for recordings with a higher fraction of periods of activity. Furthermore, for very low-noise recordings, a low percentile value yields the most accurate value for F0. For recordings that are dominated by shot noise, on the other hand, higher percentile values are more accurate, and for noise levels that are even more prominent than the true signals, a percentile value closer to the median is again the theoretically best choice. Briefly, for higher activity and low noise (seen relative to each other), you can safely reduce the percentile, while it needs to be increased towards the median for lower activity levels and high noise levels. In practice, it is useful to plot the computed F0 together with the recorded fluorescence trace for typical recordings and visually judge whether the F0 value is indeed a good estimate for the baseline fluorescence or not.
Computing F0 using a moving percentile
Next, it is also possible to compute moving percentiles or moving medians instead of simple percentiles or medians. The idea behind this approach is that there are slow drifts of the baseline F0 that should be accounted for by slowly adjusting F0. This approach is currently the most commonly used one in source extraction packages like CaImAn or Suite2p, since it does not harm in most cases and can rescue a drift from contaminating all analyses. Such a drift may occur due to several reasons. First, due to bleaching of the fluorescent indicator, an effect that is not desirable because bleaching also decreases the signal-to-noise ratio within the same recording. In this case, normalizing by an exponential or linear fit may be a good way to compensate for this drift. Alternatively, such a drift can also be due to a drift of the observed neurons out of the microscope’s focal spot due to a minor instability of a head-fixation apparatus or due to warming up of critical microscopy components that determine the focus of the scanning beam such as resonant scanners or tunable lenses. Ideally, such a focus shift should however been checked for in the raw data and be prevented during experiments. If such small drifts do occur in existing datasets – and they may also occur without you fully understanding them – moving percentiles are a good way to correct for them, and most standard toolboxes (Suite2p, CaImAn, EXTRACT) use them. That’s how such a code could look like.
Matlab:
% Parameters w_size = 600; % Moving window size q_value = 20; % 20th percentile x_w = floor(w_size/2); % helper variable % Apply moving percentile filter F0 = arrayfun(@(x) prctile(fluo_trace(max(1,x-x_w):min(end,x+x_w)), q_value), 1:length(fluo_trace));
Python:
# Import dependencies import numpy as np from scipy.ndimage import generic_filter # Parameters window_size = 600 # Moving window size (20 s for 30 frames/s) percentile_value = 20 # 20th percentile # Apply moving percentile filter F0 = generic_filter(fluorescence_trace, function=lambda x: np.percentile(x, percentile_value), size=window_size)
The typical time window used for these moving windows is something like 20 or 60 seconds – much slower than the large majority of calcium transients observed in cortical pyramidal neurons. However, in other CNS regions (e.g., spinal cord), in other neuronal types (e.g., fast-spiking interneurons), and in other cell types (e.g., astrocytes), calcium signals or the underlying spiking activity may change on a much slower timescale. This is also often true for the signals recorded from neuromodulators such as noradrenaline or acetylcholine. In this case, the moving filter should be carefully adapted in order not to accidentally remove slowly changing biological signals.

Calcium imaging of different cell types in hippocampal CA1 in behaving mice. The activity of pyramidal neurons is sparse, such that the moving window for computation of F0 does not need to exceed 10-20 s. For the selected interneuron or astrocyte, a moving window of 20 s would subtract slowly evolving true calcium changes, and a much longer time window would to compute the moving average F0 is necessary.. Data acquired by Sian Duss and Peter Rupprecht.
In cases where both slowly changing biological signals and drift e.g. due to bleaching occur at the same time, conclusions about observations have to made very carefully. Or, to put it more directly, if you see slowly changing neuronal activity, you have to convince yourself ten times before you can state that the slow changes are really of biological origin!
Computing F0 from a pre-stimulus time window
Finally, it is also common practice to compute F0 not for an entire recording but based on a short time window before a time window of interest (e.g., a repeated behavioral trial start, or a repeated sensory stimulation). This normalization based on the pre-stimulus window may be used to remove any kind of drift systematically. The crucial point is: whether such a trial-based F0 baseline computation or another definition of F0 is better depends on the biological questions asked with this analysis. In one case (pre-stimulus window normalization), the focus is on measuring the change of activity compared to the pre-stimulus window. In the other case (global normalization), the focus is on measuring the change of activity compared to a more global baseline, and it is then possible to more directly compare stimulation windows across repetitions.
Ultimately, there is no one-size-fits-all approach to computing F0 – the choice depends on noise characteristics, activity levels, and cell types, but also on the biological questions that are being addressed.
How to interpret ∆F/F
What does a certain ∆F/F mean?
The most difficult part of analyzing ∆F/F is its interpretation. The central question is: What does a certain ∆F/F transient mean? Does it reflect a single spike, multiple spikes, or a long burst of spikes? The short answer: it depends. The long answer: it depends on the cell type, the brain region, the calcium indicator, and possibly on other factors. In practice, and especially for indicators like GCaMP6f, an isolated calcium transient in a cortical pyramidal neuron is due to a burst of spikes, not a single isolated spike.
Ground truth recordings to understand ∆F/F
To get an intuition for the relationship between spikes and calcium transients, I’d recommend looking at data where this has been measured directly (so-called “ground truth” recordings). Adrian Roggenbach and I been setting up such a browsable database online, and feel free to use or modify it for your purposes!
First of all, it is important to understand that a transient with a peak ∆F/F amplitude of 30% can be due to a single spike in one neuron and due to two spikes in another one, that this may differ also between cell types and calcium indicators. Here are a few rules of thumb:
- For the commonly used GCaMP6, the smallest visible transients are often not reflecting single spikes but rather bursts. This is due to the non-linearity and kD values of GCaMP6. For GCaMP8m and GCaMP8s, the smallest visible transients are more often due to single spikes.
- Direct comparisons of ∆F/F values between pairs of neurons are problematic because resting calcium concentrations or cell-intrinsic buffering may vary from cell to cell. Furthermore, F0 is difficult to determine for GCaMP indicators due to the low baseline fluorescence.
- A single spike can trigger a calcium transient with a peak amplitude of typically 10-50% for GCaMP6 or 30-80% for GCaMP8. This is only an order of magnitude because it depends on cell types, the variants of GCaMP, and hinges on the determination of F0, which is – as discussed above – difficult.
- The values discussed so far apply to cortical pyramidal neurons (layer 2/3). For different cell types like fast-spiking interneurons, the ∆F/F transient evoked by a single spike is barely 1/10th of that value. A prominent calcium transient in a fast-spiking interneuron should therefore be interpreted as a reflection of easily 10-30 spikes, and single spikes cannot be resolved due to their small amplitude with standard population imaging. That’s why performing such ground truth across cell types and brain regions is crucial.
Spike inference from ∆F/F data
Beyond this rather qualitative interpretation of ∆F/F, it is also possible to use algorithms for spike inference, which often make an estimate of absolute spike rates. I’ve worked myself on such an algorithm based on deep learning (CASCADE), but there are also other algorithms that use a modeling approach (e.g., MLSpike). Both algorithms have in common that they are calibrated and optimized by ground truth recordings, where both calcium and electrophysiological signals are recorded from the same neuron. The inferred spike rates that you get with a very good calibrated algorithm still has to be taken as an estimate with an uncertainty. From my experience, the inferred spike rate may be off by a factor of 2, i.e., being between 50% or 200% of the true estimate. There may be an additional bias when applying spike inference on brain regions, calcium indicators or other conditions that have not been optimized for with the respective algorithm.
Then, some approaches for spike inference like the standard ones used by CaImAn or Suite2p, do not make a quantitative estimate of spike rates and are therefore at least not wrong. Personally, however, I’m convinced that it makes sense to push for quantitative and calibrated spike inference, in order to make calcium imaging more interpretable in the future. This is an active area of future research that I find really exciting. It will require not only more linear calcium indicators and a better understanding of the variability of calcium responses across neurons, but also even better algorithms.
Common pitfalls in interpretation
I will only briefly touch upon several other aspects that need to be considered when interpreting ∆F/F data. For example, there are many reports of negative deflections of ∆F/F values in the literature, often interpreted as inhibition. In many cases, these may also be brain motion artifacts or artifacts due to analysis. Negative deflections are in principle possible when the baseline firing rate is not zero, but there are two observations suggesting that negative deflections are artifacts: first, if the onset time of negative deflections is equally fast as for positive deflections; this would not be expected due to the slower off-kinetics of calcium indicators. Second, if there is a roughly equal amount of positively and negatively modulated neurons, indicating that half of the neurons are jumping out of the focus and the other half are jumping into the focus during brain motion.
Neuropil are the often unresolved arborizations of neurons that generate a blurry background. The time trace of a specific neuron can therefore be contaminated by this background. This unmixing of neuronal activity and background is difficult by principle because the activity of the two might be correlated. Depending on the brain region, activity patterns and labeling density, one may just ignore the problem, or use a linear subtraction of a neuropil background component (done by toolboxes like Suite2p, CaImAn or FISSA). An alternative approach is to use only somatic expression of the calcium indicator, preventing the bleedthrough experimentally.
Finally, there is no measurement of a thing without perturbing it at the same time. There are multiple ways how the expression of calcium indicators can perturb natural activity patterns, resulting in some cases even in epileptic or other aberrant activity patterns. These effects are not really understood yet. What is commonly accepted, however, is that overexpression and the typical “nuclear filling” with calcium indicators is a bad sign for neuronal health. High expression levels also affect calcium dynamics directly because these overly expressed calcium indicators bind calcium ions as buffers and thereby slow down the apparent calcium dynamics. That is, if you see very slow and long transients, you may have overexpressed your calcium indicator.
Final thoughts
Overall, as with any technique, calcium imaging comes with limitations and caveats. Any good experimenter or data analyst should be aware of these and keep learning about them. Because only when we understand our tools and their limitations, we can use them with confidence and interpret the biological results with precision. From my own perspective, even after working with calcium imaging in various species, brain regions, cell types, and calcium indicators for more than ten years, I still learn something new about it every now and then.
About the author
Peter Rupprecht is a junior group leader at the University of Zurich. He is studying calcium signals and their relationship to action potentials and plasticity in neurons, as well as calcium signals in astrocytes. On the side, Peter is running a blog on neuroscience and neurotechnology. You can read his most recent work on bioRxiv: Spike inference from calcium imaging data acquired with GCaMP8 indicators.

Peter Rupprecht, University of Zurich






)